Supercoiled pHOT-1 DNA is the Perfect Substrate for Assaying any Topoisomerase Enzyme
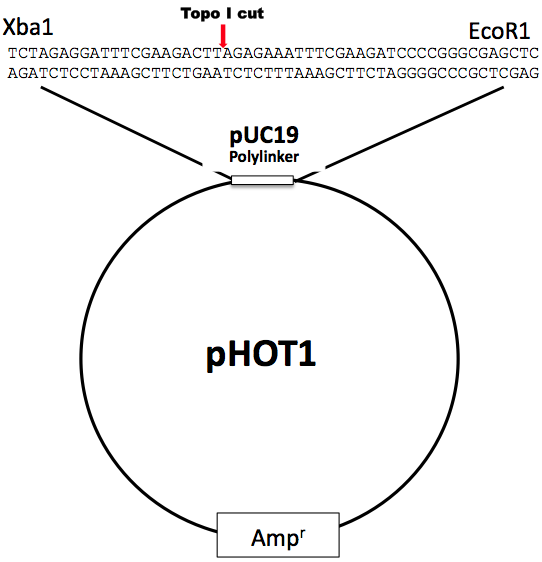
pHOT-1 is relatively small (<3KB) which is ideal for displaying intermediate topoisomers. Since all topoisomerases display degenerate DNA binding and cleavage sequences, pHOT-1 will effectively support activity for a wide array of topoisomerase. Supercoiled pHOT-1 is a scaled-down derivative of pBR322 and optimized for higher yields. It has one additional feature for researchers who wish to assess human topo I cleavage at a single sequence: we have included the human topoisomerase I preferential binding site (the hexadecameric sequence described by Westergaard and co-workers, see Bonven et al., 1985) that is derived from the tetrahymena ribosomal gene repeat. This construct is useful as a topo I cleavage substrate since the enzyme efficiently cleaves at the hexadecameric site in the absence of camptothecin. This allows the investigator to rigorously evaluate new potential topo I inhibitors because a single background cleavage is a built-in control; if the agent stimulates cleavage at this and/or any other sequence in the fragment that contains the hexadecameric site, prominent cutting is readily revealed. Typically, it is difficult to trap topo I cleavages on DNA; one must use more enzyme since cleavage complexes consume the enzyme stoichiometrically. For this reason, higher levels of enzyme are required for analysis of DNA cleavages. The fragment can be isolated from the polylinker using the sites shown below (protocols in Maniatis cloning book), end labeled and used as a cleavage substrate.
In addition, this plasmid works as well or better than pBR322 for all topoisomerases in our inventory. The small size of pHOT-1 also makes it easy to display the Gaussian distribution of topoisomers that often result from topoisomerase activity and is diagnostic for the enzyme class.

